The goal of tidysummary is to streamlines the analysis of clinical data by automatically selecting appropriate statistical descriptions and inference methods based on variable types. See the vignette for more details.
You can install the development version of tidysummary like so:
if (!requireNamespace("remotes", quietly = TRUE)) {
install.packages("remotes")
}
remotes::install_github("htqqdd/tidysummary")library(tidysummary)
result <- iris %>%
add_var() %>%
add_summary() %>%
add_p()#Here is an prepared dataset
iris <- iris %>%
mutate(group = factor(rep(1:3, each = 50),
labels = c("group1", "group2", "group3")))
#Now use tidysummary
library(tidysummary)
result <- iris %>%
add_var() %>%
add_summary(binary_show = "all") %>%
add_p()View(result)kableExtra or others your
prefer)library(kableExtra)
result[is.na(result)] <- ""
result %>%
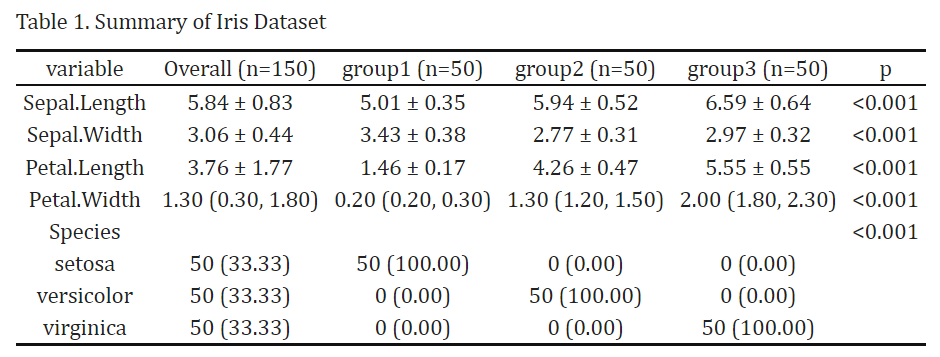
kbl(caption = "Table 1. Summary of Iris Dataset",
row.names = F,
align = "c") %>%
kable_classic(full_width = FALSE, html_font = "Cambria")
result %>%
writexl::write_xlsx("./test.xlsx")
Need a high-speed mirror for your open-source project?
Contact our mirror admin team at info@clientvps.com.
This archive is provided as a free public service to the community.
Proudly supported by infrastructure from VPSPulse , RxServers , BuyNumber , UnitVPS , OffshoreName and secure payment technology by ArionPay.