The goal of tidyheatmaps is to simplify the generation of publication-ready heatmaps from tidy data. By offering an interface to the powerful pheatmap package, it allows for the effortless creation of intricate heatmaps with minimal code. By offering an interface to the powerful pheatmap package, it allows for the effortless creation of intricate heatmaps with minimal code.
You can install the released version of tidyheatmaps from CRAN with:
install.packages("tidyheatmaps")And the development version from GitHub with:
# install.packages("devtools")
devtools::install_github("jbengler/tidyheatmaps")Given a tidy data frame of gene expression data like
data_exprs, you can easily generate a customized heatmap.
The full documentation can be found here.
library(tidyheatmaps)
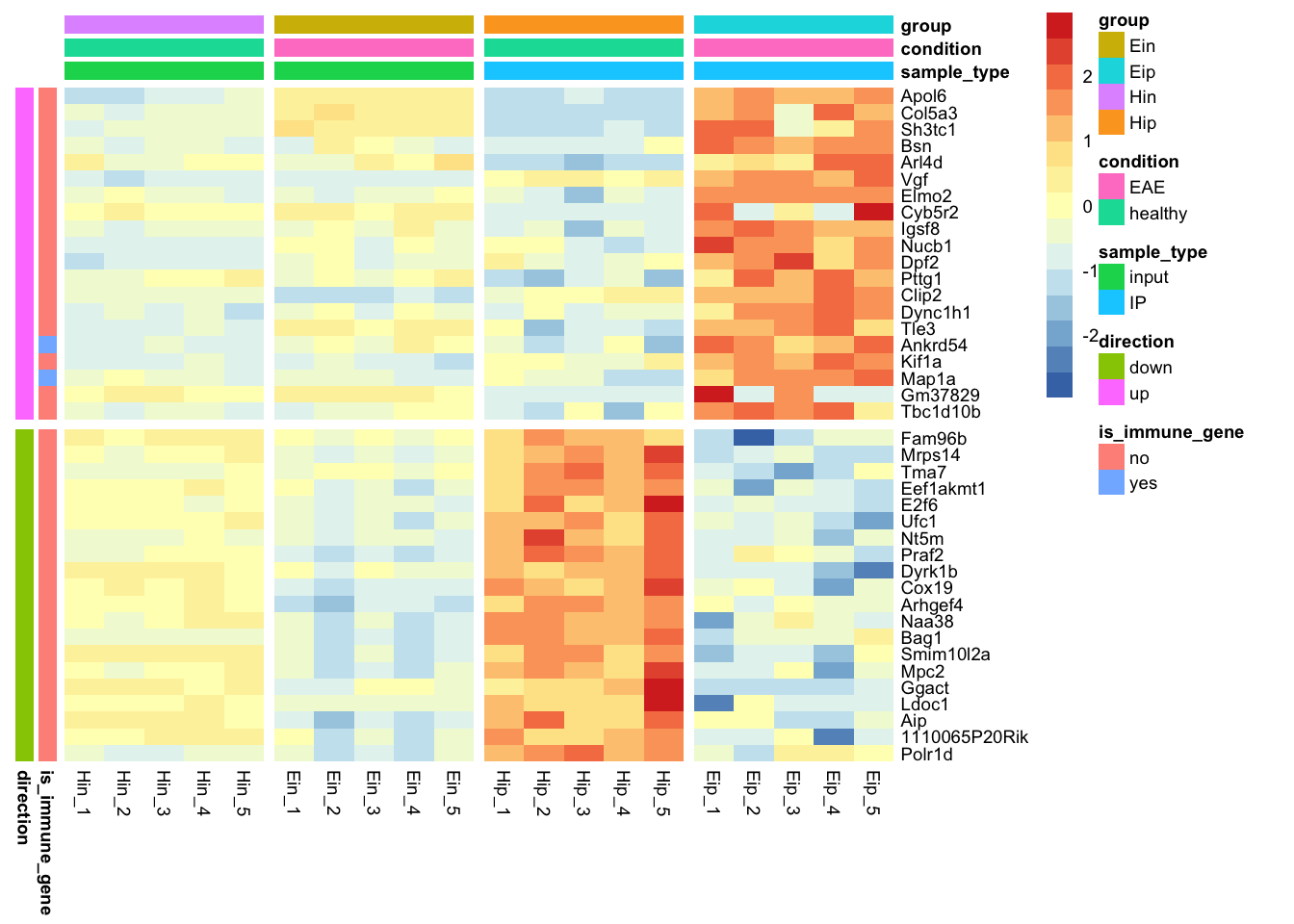
tidyheatmap(data_exprs,
rows = external_gene_name,
columns = sample,
values = expression,
scale = "row",
annotation_col = c(sample_type, condition, group),
annotation_row = c(is_immune_gene, direction),
gaps_row = direction,
gaps_col = group
)
tidyplots relies on a number of fantastic packages that do all the heavy lifting behind the scenes. These include dplyr, pheatmap, rlang, grDevices, tidyr, tibble and RColorBrewer.
Need a high-speed mirror for your open-source project?
Contact our mirror admin team at info@clientvps.com.
This archive is provided as a free public service to the community.
Proudly supported by infrastructure from VPSPulse , RxServers , BuyNumber , UnitVPS , OffshoreName and secure payment technology by ArionPay.