Fully Latent Principal Stratification (FLPS) is an extension of principal stratification.
Install the latest release from CRAN or git repository:
devtools::install_github("sooyongl/flps")
install.packages("flps")library(flps)## Version: 1.0.0
##
## It is a demo.
## Acknowledgements. It is supported by the Institute of Education Sciences, U.S. Department of Education, through Grant R305D210036.binary: a data frame containing all the data for FLPS.
It is used in runFLPS function.rstan
package.data(binary)# Input data frame
data.table::data.table(binary)## schid id sex race pretest stdscore cm_sex cm_race cm_pretest
## <int> <int> <int> <int> <int> <num> <num> <num> <num>
## 1: 1 2383 0 1 20 -0.3296 0.4406780 0.9322034 14.81356
## 2: 1 2384 1 0 8 1.1597 0.4406780 0.9322034 14.81356
## 3: 1 2385 0 1 14 -0.7385 0.4406780 0.9322034 14.81356
## 4: 1 2387 0 1 12 -1.3518 0.4406780 0.9322034 14.81356
## 5: 1 2388 0 1 6 -1.2057 0.4406780 0.9322034 14.81356
## ---
## 4762: 63 4761 0 1 16 0.6457 0.4860558 0.3944223 17.03187
## 4763: 63 4763 0 0 11 0.3168 0.4860558 0.3944223 17.03187
## 4764: 63 4764 0 0 18 -0.4114 0.4860558 0.3944223 17.03187
## 4765: 63 4765 1 0 31 2.2116 0.4860558 0.3944223 17.03187
## 4766: 63 4766 1 1 13 -0.0668 0.4860558 0.3944223 17.03187
## cm_stdscore trt Y q1 q2 q3 q4 q5 q6 q7 q8
## <num> <int> <num> <int> <int> <int> <int> <int> <int> <int> <int>
## 1: -0.4068119 1 -0.487 0 NA 1 1 1 1 1 NA
## 2: -0.4068119 1 0.487 1 NA 1 1 1 1 1 1
## 3: -0.4068119 1 -1.073 0 NA 1 1 0 NA NA NA
## 4: -0.4068119 1 -1.087 0 NA 1 1 1 NA NA NA
## 5: -0.4068119 1 -1.171 0 NA 1 1 NA NA NA NA
## ---
## 4762: 0.0997502 0 -0.114 0 NA NA NA NA NA NA NA
## 4763: 0.0997502 0 -0.438 0 NA NA NA NA NA NA NA
## 4764: 0.0997502 0 -1.047 0 NA NA NA NA NA NA NA
## 4765: 0.0997502 0 0.765 0 NA NA NA NA NA NA NA
## 4766: 0.0997502 0 0.539 0 NA NA NA NA NA NA NA
## q9 q10 q11 q12 q13 q14 q15 q16 q17 q18 q19 q20
## <int> <int> <int> <int> <int> <int> <int> <int> <int> <int> <int> <int>
## 1: NA NA NA NA NA NA NA NA NA NA NA NA
## 2: 1 1 1 1 NA NA NA NA NA NA NA NA
## 3: NA NA NA NA NA NA NA NA NA NA NA NA
## 4: NA NA NA NA NA NA NA NA NA NA NA NA
## 5: NA NA NA NA NA NA NA NA NA NA NA NA
## ---
## 4762: NA NA NA NA NA NA NA NA NA NA NA NA
## 4763: NA NA NA NA NA NA NA NA NA NA NA NA
## 4764: NA NA NA NA NA NA NA NA NA NA NA NA
## 4765: NA NA NA NA NA NA NA NA NA NA NA NA
## 4766: NA NA NA NA NA NA NA NA NA NA NA NArunFLPS() internally transforms binary data into the
format suitable for rstan, subsequently executing
FLPS.
To avoid re-compiling the Stan code each time, pre-compile it
using modelBuilder(), which stores the stanmodel object in
the flps directory, accelerating subsequent
analyses.
Once the Stan model is compiled, use importModel()
to bring in the compiled Stan code. This code can then be provided to
the compiled_stan argument in runFLPS. If this
step is omitted, runFLPS() will compile the Stan code
during each execution of FLPS.
modelBuilder(type = "rasch")
complied_stan <- importModel(type = "rasch")rstan and
StanHeaders packages.remove.packages(c("rstan", "StanHeaders"))
install.packages("rstan", repos = c("https://mc-stan.org/r-packages/", getOption("repos")))Now, execute your FLPS model. Given the time-intensive nature of the process, chains and iterations have been initially limited to 1 and 5000, respectively. It is advisable to increase these values for your specific research needs.
# Subset of data: 1000 students
binary <- binary[c(sample(which(binary$trt == 1), 250),
sample(which(binary$trt == 0), 250)),]
res <- runFLPS(
inp_data = binary,
# complied_stan = complied # if necessary
outcome = "Y",
trt = "trt",
covariate = c("sex","race","pretest","stdscore"),
lv_type = "rasch",
lv_model = "F =~ q1 + q2 + q3 + q4 + q5 + q6 + q7 + q8 + q9 + q10",
stan_options = list(iter = 5000, cores = 1, chains = 2)
)Retrieve summaries and visualize results with the following:
summary(res)The flps_plot() shows the plot related to FLPS
models
flps_plot(res, type = "causal")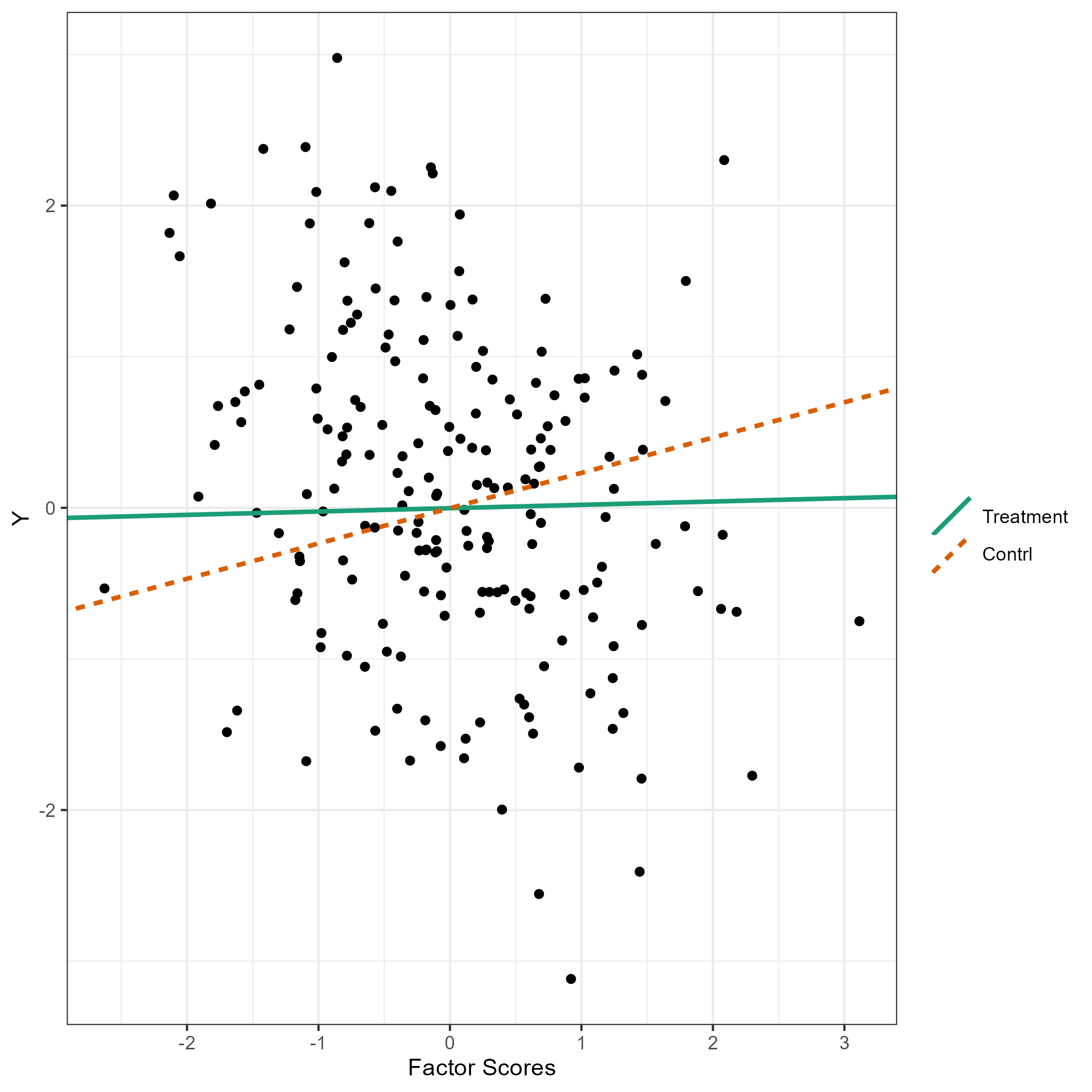
flps_plot(res, type = "latent")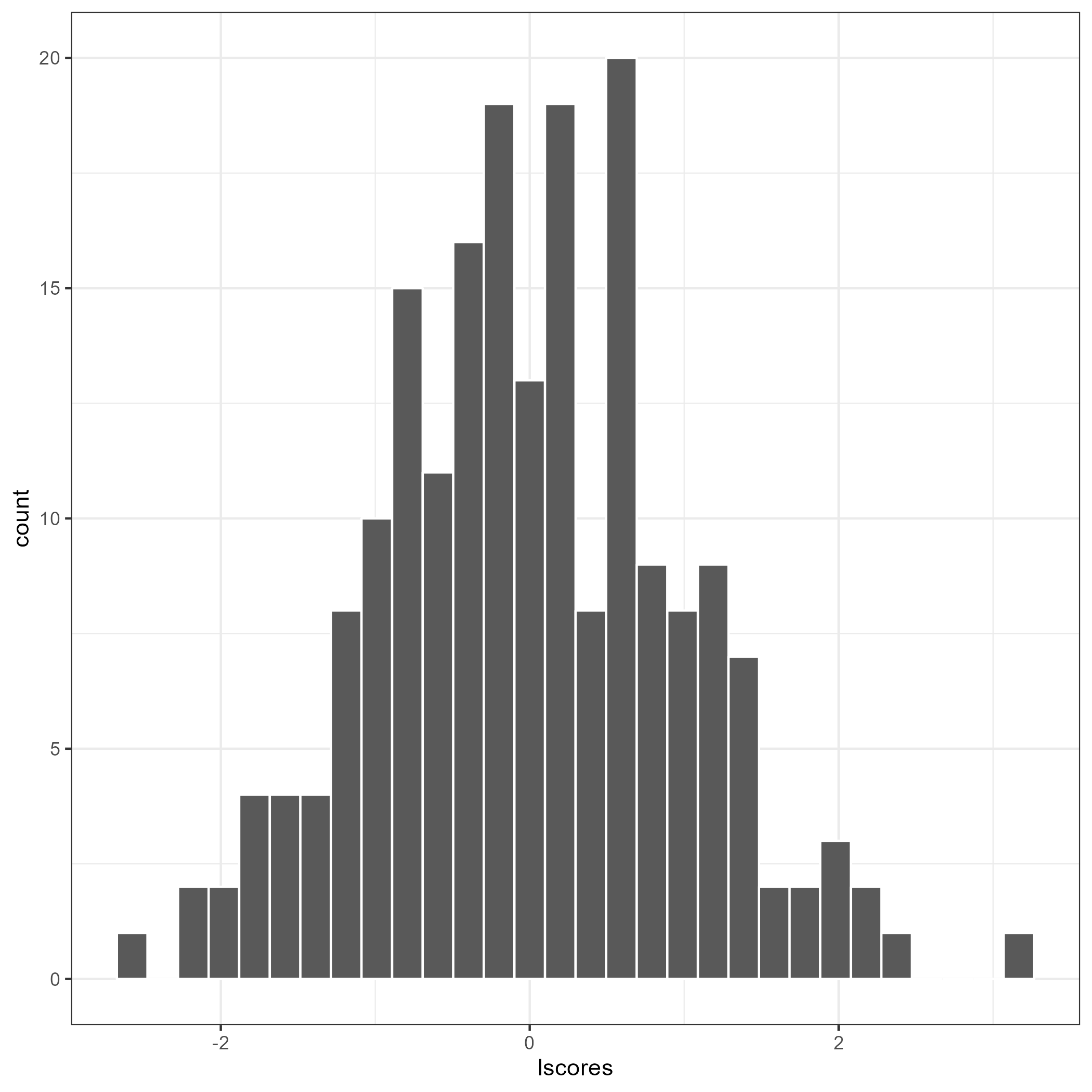
Acknowledgements. This package is supported by the Institute of Education Sciences, U.S. Department of Education, through Grant R305D210036.↩︎
Need a high-speed mirror for your open-source project?
Contact our mirror admin team at info@clientvps.com.
This archive is provided as a free public service to the community.
Proudly supported by infrastructure from VPSPulse , RxServers , BuyNumber , UnitVPS , OffshoreName and secure payment technology by ArionPay.