BayesMallowsSMC2 provides functions for performing sequential inference in the Bayesian Mallows model using the SMC2 algorithm (Sørensen et al. 2025).
You can install the released version of BayesMallowsSMC2 from CRAN with:
install.packages("BayesMallowsSMC2")Or install the development version of BayesMallowsSMC2 from GitHub with:
# install.packages("devtools")
devtools::install_github("osorensen/BayesMallowsSMC2")Here is a basic example using the included
complete_rankings dataset:
library(BayesMallowsSMC2)
# Fit the model with complete rankings
set.seed(123)
mod <- compute_sequentially(
complete_rankings,
hyperparameters = set_hyperparameters(n_items = 5),
smc_options = set_smc_options(n_particles = 100, n_particle_filters = 1)
)
# Show model info
mod
#> BayesMallowsSMC2 Model
#> ======================
#>
#> Number of particles: 100
#> Number of timepoints: 100
#> Number of items: 5
#> Number of clusters: 1
#>
#> Log marginal likelihood: -472.34
#> Final ESS: 63.55
#> Resampling events: 5/100We can visualize the posterior distributions of the parameters:
# Posterior distribution of alpha (dispersion parameter)
plot(mod, parameter = "alpha")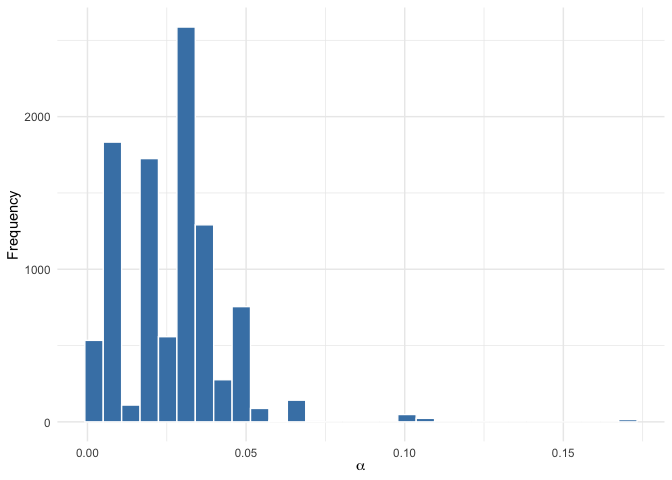
# Posterior distribution of rho (ranking positions)
plot(mod, parameter = "rho", items = c(1, 2, 3))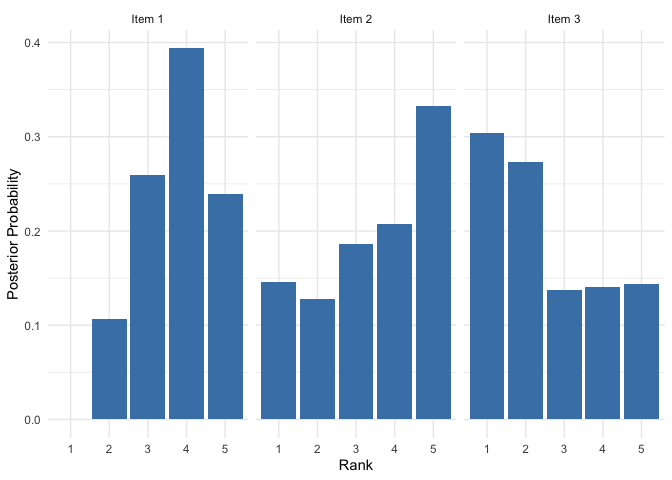
Need a high-speed mirror for your open-source project?
Contact our mirror admin team at info@clientvps.com.
This archive is provided as a free public service to the community.
Proudly supported by infrastructure from VPSPulse , RxServers , BuyNumber , UnitVPS , OffshoreName and secure payment technology by ArionPay.